Free Access
| Issue |
Genet. Sel. Evol.
Volume 39, Number 3, May-June 2007
|
|
|---|---|---|
| Page(s) | 301 - 317 | |
| DOI | https://doi.org/10.1051/gse:2007005 | |
| Published online | 14 April 2007 | |
References of
Genet. Sel. Evol. 39 (2007) 301-317
- Bennewitz J., Reinsch N., Reinhart F., Liu Z., Kalm E., Top down preselection using marker assisted estimates of breeding values in dairy cattle, J. Anim. Breed. Genet. 121 (2004) 307-318.
- Boichard D., Fritz A., Rossignol M.N., Boscher M.Y., Malafosse A.,
Colleau J.J., Implementation of marker-assisted selection in French dairy
cattle. Proceedings of 7
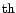 World Congress on Genetics Applied to
Livestock Production, 19-23 August 2002, Montpellier, France, CD
communication no. 22-03.
World Congress on Genetics Applied to
Livestock Production, 19-23 August 2002, Montpellier, France, CD
communication no. 22-03.
- Coimbra M.R.M., Kabayashi K., Koretsugu S., Hasegawa O., Ohara E., Ozaki S., Sakamoto T., Naruse K., Okamoto N., A genetic map of the Japanese flounder Paralichthys olivaceus, Aquaculture 220 (2003) 203-218 [CrossRef].
- Dekkers J.C.M, Hospital F., The use of molecular genetics in the improvement of agricultural populations, Nat. Genet. Rev. 3 (2002) 22-32.
- Doyle R.W., Herbinger C.M., The use of DNA fingerprinting for
high-intensity within-family selection in fish breeding, Proceedings of the
5
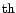 World Congress on Genetics Applied to Livestock Production, August
7-12 1994, Vol. 19, Guelph, Canada, pp. 364-371.
World Congress on Genetics Applied to Livestock Production, August
7-12 1994, Vol. 19, Guelph, Canada, pp. 364-371.
- Falconer D.S., Mackay T.F.C., Introduction to quantitative genetics,
4
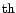 edn., Longman Sci. and Tech., Harlow, UK, 1996.
edn., Longman Sci. and Tech., Harlow, UK, 1996.
- Fernando R.L., Grossman M., Marker-assisted selection using best linear biased prediction, Genet. Sel. Evol. 21 (1989) 467-477.
- Gibson J.P., Short term gain at the expense of long term response with
selection of identified loci. Proceedings of the 5
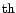 World Congress on
Genetics Applied to Livestock Production, August 7-12 1994, Vol. 21, Guelph,
Canada, pp. 201-204.
World Congress on
Genetics Applied to Livestock Production, August 7-12 1994, Vol. 21, Guelph,
Canada, pp. 201-204.
- Gjedrem T., Olesen I., Basic statistical parameters, in: Gjedrem T. (Ed.), Selection and breeding programs in aquaculture, Springer, Doordrecht, the Netherlands, 2005, pp. 45-72.
- Goddard M.E., A mixed model analyses of data on multiple genetic markers, Theor. Appl. Genet. 83 (1992) 878-886 [CrossRef].
- Kashi Y., Hallerman E., Soller M., Marker assisted selection of candidate bulls for progeny testing programs, Anim. Prod. 51 (1990) 63-74.
- Lande R., Thompson R., Efficiency of marker-assisted selection in the improvement of quantitative traits, Genetics 124 (1990) 743-756 [PubMed].
- Lee B.Y., Lee W.J., Streelman T., Carleton K.L., Howe A.E., Hulata G., Slettan A., Stern J.E., Terai Y., Kocher T.D., A second-generation genetic linkage map of tilapia (Oreochromis spp.), Genetics 170 (2005) 237-244 [CrossRef] [PubMed].
- Mackinnon M.J., Georges M.A.J., Marker-assisted preselection of young dairy sires prior to progeny-testing, Livest. Prod. Sci. 54 (1998) 229-250 [CrossRef].
- Martyniuk C.J., Perry G.M.L., Mogahadam H.K., Ferguson M.M., Danzmann R.G., The genetic architecture of correlations among growth-related traits and male age at maturation in rainbow trout, J. Fish Biol. 63 (2003) 746-764.
- Meuwissen T.H.E., Maximizing the response of selection with a predefined rate of inbreeding, J. Anim. Sci. 75 (1997) 934-940 [PubMed].
- Meuwissen T.H.E., Goddard M.E., The use of marker haplotypes in animal breeding schemes, Genet. Sel. Evol. 28 (1996) 161-176.
- Meuwissen T.H.E., Sonesson A.K., Genotype-assisted optimum contribution selection to maximise selection response over a specified time period, Genet. Res. 84 (2004) 109-116 [CrossRef] [PubMed].
- Moen T., Hoyheim B., Munck H., Gomez-Raya L., A linkage map of Atlantic salmon (Salmo salar) reveals an uncommonly large difference in recombination rate between the sexes, Anim. Genet. 35 (2004) 81-92 [PubMed].
- Moen T., Sonesson A.K., Hayes B.J., Lien S., Munck H., Meuwissen T.H.E., Mapping of a quantitative trait loci for resistance against infectious salmon anemia in Atlantic salmon (Salmo salar): comparing survival models with binary models, BMC (2006) Submitted.
- Nichols K.M., Young W.P., Danzmann R.G., Robison B.D., Rexroad C., Noakes M., Phillips R.B., Bentzen P., Spies I., Knudsen K., Allendorf F.W., Cunningham B.M., Brunelli J., Zhang H., Ristow S., Drew R., Brown K.H., Wheeler P.A., Thorgaard G.H., A consolidated linkage map for rainbow trout (Oncorhynchus mykiss), Anim. Genet. 34 (2003a) 102-115 [PubMed].
- Nichols K.M., Bartholomew J., Thorgaard G.H., Mapping multiple genetic loci associated with Ceratomyxa shasta resistance in Oncorhynchus mykiss, Diseas. Aq. Org. 56 (2003b) 145-154.
- O'Malley K.G., Sakamoto T., Danzmann R.G., Ferguson M.M., Quantitative trait loci for spawning date and body weight in rainbow trout: testing for conserved effects across ancestrally duplicated chromosomes, J. Hered. 94 (2003) 273-284 [CrossRef] [PubMed].
- Ozaki A., Sakamoto T., Khoo S., Nakamura K., Coimbra M.R.M., Akutsu T., Okamoto N., Quantitative trait loci (QTLs) associated with resistance/susceptibility to infectious pancreatic necrosis virus (IPNV) in rainbow trout (Oncorhynchus mykiss), Mol. Genet. Genom. 265 (2001) 23-31.
- Reid D.P., Szanto A., Glebe B., Danzmann R.G., Ferguson M.M., QTL for body weight and condition factor in Atlantic salmon (Salmo salar): comparative analysis with rainbow trout (Oncorhynchus mykiss) and Arctic charr (Salvelinus alpinus), Heredity 94 (2005) 166-172 [CrossRef] [PubMed].
- Rodriguez M.F., LaPatra S., Williams S., Famula T., May B., Genetic markers associated with resistance to infectious hematopoietic necrosis in rainbow and steelhead trout (Oncorhynchus mykiss) backcrosses, Aquaculture 241 (2005) 93-115 [CrossRef].
- Sakamoto T., Danzmann R.G., Gharbi K., Howard P., Ozaki A., Khoo S.K., Woram R.A., Okamoto N., Ferguson M.M., Holm L.-E., Guyomard R., Hoyheim B., A microsatellite linkage map of rainbow trout (Oncorhynchus mykiss) characterized by large sex-specific differences in recombination rates, Genetics 155 (2000) 1331-1345 [PubMed].
- Sonesson A.K., A combination of walk-back and optimum contribution selection for fish, Genet. Sel. Evol. 37 (2005) 587-599 [EDP Sciences] [CrossRef] [PubMed].
- Villanueva B., Pong-Wong R., Woolliams J.A., Marker assisted selection with optimised contributions of the candidates to selection, Genet. Sel. Evol. 34 (2002) 679-703 [EDP Sciences] [CrossRef] [PubMed].
- Weller J.I., Kashi Y., Soller M., Power of daughter and granddaughter designs for determining linkage between marker loci and quantitative trait loci in dairy cattle, J. Dairy Sci. 73 (1990) 2525-2537 [PubMed].
- Woram R.A., McGowan C., Stout J.A., Gharbi K., Ferguson M.M., Hoyheim B., Davidson E.A., Davidson W.S., Rexroad C., Danzmann R.G., A genetic linkage map for Arctic char (Salvelinus alpinus): evidence for higher recombination rates and segregating distortion in hybrid versus pure strain mapping parents, Genome 47 (2004) 304-315 [PubMed].


